NUS Reconstruction of HNCA with Virtual Decoupling in SMILE
National Institute of Diabetes and Digestive and Kidney Diseases
National Institutes of Health
Bethesda, MD 20892, U.S.A.
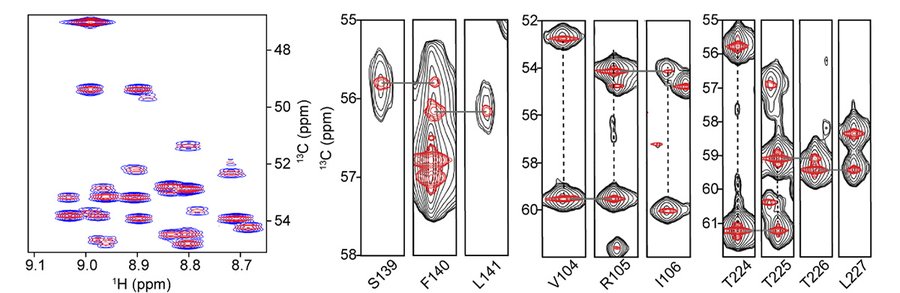
HNCA is one of the first triple resonance NMR experiments developed for studies of isotopically enriched proteins. A high quality well-resolved HNCA spectrum can greatly facilitate the resonance assignment process. However, the spectral resolution in the 13C indirect dimension is largely limited by the 35 ± 2.5 Hz 1JCC coupling. Others have developed spectral deconvolution methods, including a deep learning algorithm, to remove this coupling during NUS reconstruction. Here, we implement a general approach to virtual decoupling within the SMILE reconstruction program. When applied to the HNCA on a 68 kDa dimer of the SARS-CoV-2 main protease (MPro), our method is demonstrated to significantly improve the resolution of the experiment. Details of the implementation and the NUS sampling requirements will be discussed. We expect the virtual decoupling approach to be applicable to other types of experiments, including those with antiphase doublets, provided the couplings are sufficiently uniform. It can also be readily implemented in NUS reconstruction programs other than SMILE.
References
- Kay LE, Ikura M, Tschudin R, Bax A. Three-dimensional triple-resonance NMR spectroscopy of isotopically enriched proteins. J Magn Reson 89, 496–514 (1990).
- Ying J, Delaglio F, Torchia DA, Bax A. Sparse multidimensional iterative lineshape-enhanced (SMILE) reconstruction of both nonuniformly sampled and conventional NMR data. J Biomol NMR 68, 101-118 (2017).
- Kazimierczuk K, Kasprzak P, Georgoulia PS, Matečko-Burmann I, Burmann BM, Isaksson L, Gustavsson E, Westenhoff S, Orekhov VY. Resolution enhancement in NMR spectra by deconvolution with compressed sensing reconstruction. Chem Commun 56, 14585–14588 (2020).
- Karunanithy G, Hansen DF. FID-Net: A versatile deep neural network architecture for NMR spectral reconstruction and virtual decoupling. J Biomol NMR 75, 179–191 (2021).
Acknowledgements
This work was supported by the Intramural Research Program of the NIDDK and by the Intramural Antiviral Target Program of the Office of the Director, NIH. I would like to acknowledge Dr. Ad Bax for his support and guidance.